ASSEMBLING DISCIPLINES
EPIC Assembly project (Emergence of Novel Functional Assembly by Evo-Physico Information Coupling) explores the mechanisms and principles by which advanced and complex biological functions have emerged and been refined in biological history. We specifically focus on the concept of "assemblies", systems composed of heterotypic interacting elements, as a unifying framework that spans molecular to multicellular levels. Assemblies can become combinatorially complex as the number and diversity of constituent elements increase, offering a structural basis for the evolutionary exploration of novel functions.
To tackle this challenging problem, it is essential to integrate insights and techniques from diverse domains, ranging from biology, physics, information science, and mathematics.
In this conference, we ASSEMBLE leading researchers engaged in this cross-disciplinary field to advance the frontiers of integrative science and to explore new avenues for understanding the emergence and evolution of biological functions.
Day Event
Speakers
Participants


Antonio Celani
International Centre for Theoretical Physics
Jeremy Beaulieu
University of Arkansas
Paul François
Université de Montréal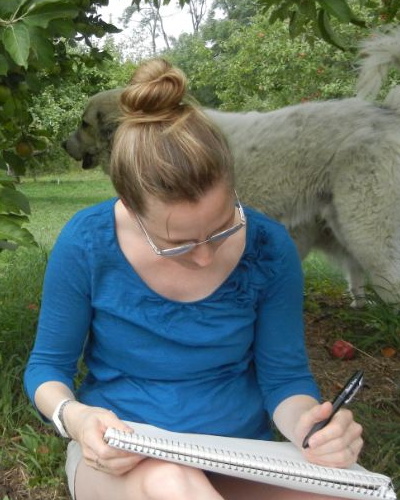
Jennifer Schwarz
Syracuse University
Hsuan-yi Chen
National Central University
Suropriya Saha
Max Planck Institute
Wenying Shou
University College London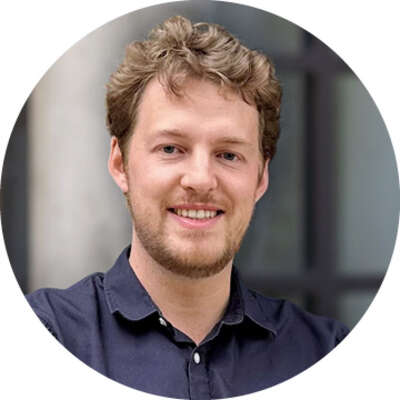
David Brückner
University of Basel
Tetsuya Hiraiwa
Institute of Physics, Academia Sinica
Lorenzo Di Michele
University of Cambridge
Yongdae Shin
Seoul National University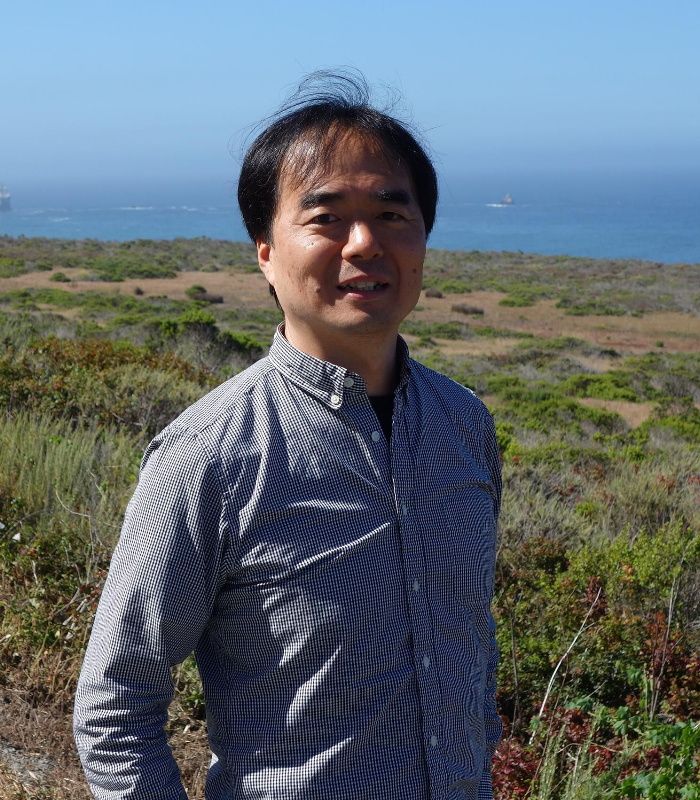
Hiroaki Matsunami
Duke University
Ikuo Masuho
Sanford Research-

Kyogo Kawaguchi
The University of Tokyo
Masahiro Takinoue
Institute of Science Tokyo
Kazuhiro AOKI
Kyoto University
Yasuka TODA
Institute of Science Tokyo
Daiki UMETSU
Osaka University
Satoshi SAWAI
University of Tokyo
Nen SAITO
Hiroshima University
Ryo HANAI
Institute of Science Tokyo
Kaoru SUGIMURA
University of Tokyo
Tetsuya J. KOBAYASHI
University of Tokyo
Takao K. SUZUKI
Juntendo University
Kazufumi HOSODA
CiNet, NICTPoster Session Times
- Mar 14, 16:45–18:30
- Mar 15, 16:45–18:30
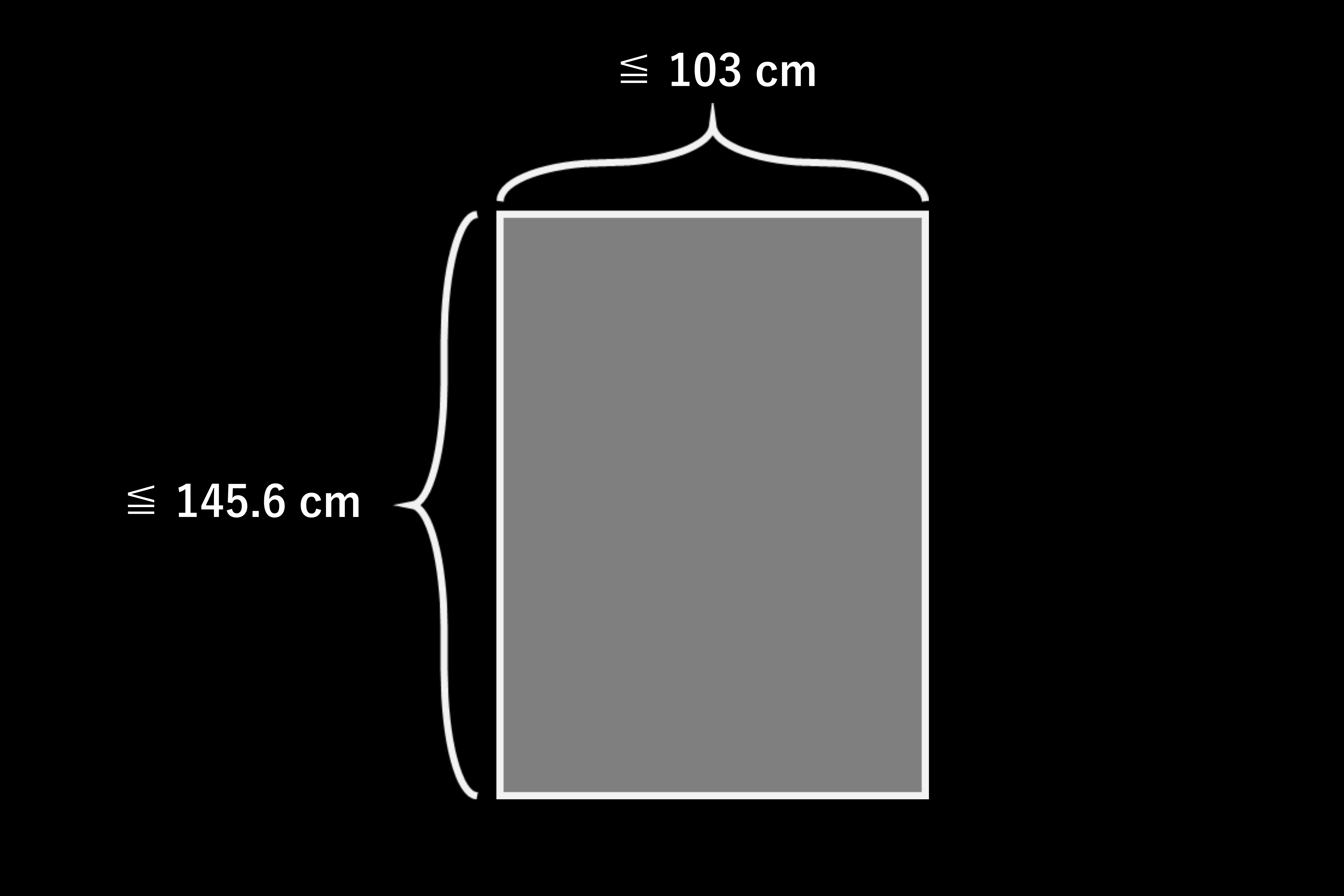
Poster size
The poster must fit within 145.6 cm (height) × 103 cm (width); standard sizes such as A0 or B0 are acceptable.
- 09:45-10:00
-
Opening remarks & EPIC assembly project - Tetsuya Kobayashi Biological systems exhibit diverse and highly specialized functions across molecular to multicellular scales. In the EPIC Assembly project, we investigate how such functions emerge and increase in complexity through evolution by focusing on “assemblies”—systems composed of heterotypic interacting elements. As the number and diversity of components grow, the space of possible assemblies expands combinatorially, raising the question of how functional configurations are selected and stabilized. We propose an idea that novel functions arise from the interplay between non-equilibrium dynamics and evolutionary processes. To test this idea, we integrate high-resolution imaging, phylogenetic reconstruction of ancestral assemblies, and their experimental reconstitution with mathematical models that describe both non-equilibrium and evolutionary dynamics. By combining experimental and theoretical approaches, we aim to identify general principles governing the emergence and evolution of functional assemblies. These insights may guide the design of synthetic biological systems and inform theoretical frameworks for complex adaptive systems.
- 10:00-10:35
-
Invited Talk: Decoding behavior with minimal and interpretable agent models - Antonio Celani Understanding how living organisms process sensory information and translate it into decisions is a fundamental problem across biological scales. We address this challenge by introducing a method to reconstruct general decision processes directly from behavioral observations alone. Our agent model is defined by a recurrent dynamics over a discrete set of internal states. We validate our method on synthetic agents and we infer agent models from experimental data of rats performing evidence accumulation and of mice making decisions under uncertainty and in changing environments. In both cases, very few internal states suffice to reproduce the observed behavior with high accuracy.
- 10:35-11:10
-
Invited Talk: Signal and Noise in Biodiversity Dynamics - Jeremy Beaulieu Across a variety of biological datasets, from genomes to conservation to the fossil record, evolutionary rates appear to increase toward the present or over short time scales. This has long been seen as an indication of processes operating differently at different time scales, even potentially as an indicator of a need for new theory connecting macroevolution and microevolution. Here I introduce a set of models that assess the relationship between rate and time and demonstrate that these patterns are statistical artifacts of time-independent errors present across ecological and evolutionary datasets, which produce hyperbolic patterns of rates through time. I will show that plotting a noisy numerator divided by time versus time leads to the observed hyperbolic pattern; in fact, randomizing the amount of change over time generates patterns functionally identical to observed patterns. Ignoring noise will not only obscure true patterns but create novel patterns that have long misled scientists. I will also briefly discuss the need for a broader term for noise (i.e., tip fog), and then present new methods for dealing with tip fog in a variety of phylogenetic comparative methods, even when the underlying variation is not actually measured.
- 11:10-11:20
-
Coffee Break
- 11:20-11:55
-
Invited Talk: Generative Epigenetic Landscape modelling - Paul François
- 11:55-12:30
-
EPIC Member Talk: Macroevolutionary Pathways of Multi-Component Phenotypic Assembly in Bacteria - Takao Suzuki How do complex phenotypes assemble over macroevolutionary time? A key conceptual framework for addressing this question is the notion of multi-component systems, which describes biological entities as structured assemblies of interacting components. This framework characterizes structural properties of assembly, yet its macroevolutionary dynamics remain poorly understood. Here, we develop a novel method to reconstruct macroevolutionary pathways of multi-component phenotypic states across nearly 4,000 bacterial species. We found that trait combinations are hierarchically structured, with motility–sporulation states forming primary evolutionary modules within which morphology diversified. We then analyzed genome-wide gene content and uncovered distinct gene repertoires underlying these hierarchical adaptive strategies. Remarkably, regulatory genes, including anti-sigma factors, coevolve with each phenotypic state transition. Together, this approach opens the door to a quantitative theory of evolutionary assembly.
- 12:30-14:00
-
Lunch Break
- 14:00-14:35
-
EPIC Member Talk: A Mathematical Framework for Deformable Cells in Multicellular Dynamics - Nen Saito Cell deformability within dense tissues can play a crucial role in determining tissue-scale mechanical properties. Simulation-based theoretical studies are essential for characterizing such collective behaviors. However, large-scale simulations of deformable cells have typically been limited to simple convex polygonal shapes due to computational constraints. In this talk, I will present a simulation framework that enables efficient modeling of large populations of freely deformable cells [1], offering significantly improved computational performance over existing methods. Using this framework, we analyze various biological phenomena, including collective dynamics in densely packed cell populations and cell assembly processes guided by chemotactic cues. (1. Saito & Ishihara (2024). Sci. Adv. 10(19), eadi8433. )
- 14:35-15:10
-
Invited Talk: Mechanochemical Collective Cellular Intelligence in Spheroids - Jennifer Schwarz In biology, learning is typically associated with brains, yet many organisms without even neurons exhibit adaptive behavior. We propose that spheroids—assemblies of mechanically coupled cells—can exhibit learning. We hypothesize that cellular deformations generate long-range mechanical fields that couple cells into distributed networks, forming associations through mechanically mediated interactions, where sequences of deformations encode learned responses, analogous to sequences of neuronal firing. Using a 3D vertex-based computational model, we test this hypothesis by demonstrating that spheroids can be trained to acquire specific cell stress patterns via mechanochemical remodeling, effectively storing collective associations. These results suggest a physical mechanism for learning as an emergent property of interacting cells and highlight general principles of intelligence applicable to biological tissues and beyond.
- 15:10-15:20
-
Coffee Break
- 15:20-15:55
-
EPIC Member Talk: Cell size control emerges from the vein-dependent coordinated divisions of distinct cell groups in Drosophila wing - Kaoru Sugimura During morphogenesis, cell divisions are precisely regulated in space and time. The biological objectives achieved by such regulation are not fully understood, particularly in tissues whose overall size is kept constant. Here, by applying a newly developed lineage-reconstruction pipeline to the Drosophila pupal wing, we reveal that the wing is composed of distinct cell groups that differ in division number, timing, and spatial positioning relative to wing veins. We show that the frequencies of these lineages, together with their initial cell sizes and growth profiles, converge to achieve a highly conserved average cell size. Our data further suggest that distance from wing veins provides spatial information that biases where distinct lineages arise, and that genetic elimination of wing veins suppresses a specific lineage and disrupts cell-size control. Finally, our results point to a multiscale organization of division patterns, in which vein-associated spatial information is integrated with local neighbor effects in a manner that would mitigate mechanical instability within the tissue. Together, these findings delineate a cell-size control mechanism based on coordinated divisions of distinct cell groups, which supports robust morphogenesis and functional tissue design [1]. In the latter part of my talk, I will also present our simulation study on intercellular force networks in soft, active, and deformable cell populations [2]. (1. Sugimura, Takayanagi, Namba, Qu, Ishihara. bioRxiv 2025; 2. Yoshida, Saito, Ishihara, Sugimura. In preparation.)
- 15:55-16:30
-
Invited Talk: How do incorrect ligands help detect a correct ligand? - Hsuan-Yi Chen I will introduce a simple statistical physical model for a cluster of receptors that can have similar properties as T cell receptors. Our numerical simulations show that a cluster of receptors that bind reversibly to two types (correct/incorrect) of ligands in the environment with binding-state coupling between nearest neighboring receptors and a simple kinetic proofreading scheme for the activation/deactivation of receptors can have specific, speedy, and strong response even when there is only one correct ligand. This is achieved by the enhanced binding and lifetime of incorrect ligand-receptor complexes in the presence of a correct one.
- 16:30-16:45
-
Coffee Break
- 16:45-18:30
-
Poster Session
- 10:00-10:35
-
EPIC Member Talk: - Ryo Hanai
- 10:35-11:10
-
Invited Talk: Pattern formation in number conserving active systems - Suropriya Saha
- 11:10-11:20
-
Coffee Break
- 11:20-11:55
-
EPIC Member Talk: Eureka, Ecosystems, and Ultra-Efficient AI Hardware: Beyond Bio- or Brain-Inspired Alone - Kazufumi Hosoda Human Eureka enables us “to solve unexpected problems.” More fundamentally, a central requirement of living systems in evolution is adaptability: “to cope with unpredictable disturbances.” We propose that Eureka-like transitions, organism-level adaptability, and ecosystem-level robustness share common information-processing principles realized by far-from-equilibrium biological assemblies. Motivated by this hypothesis, we take three complementary routes. (i) Brain (highly organized assembly): we measure moments of insight (“Eureka!”) using fMRI and model the underlying dynamics with recurrent neural networks that leverage chaotic exploration to generate insight-like dynamics. (ii) Ecosystems (fully decentralized assemblies): we construct synthetic ecosystems enabling high-throughput tests of robustness and adaptation with no central control. (iii) Hardware: we discuss ultra-energy-efficient computing architectures that harness fluctuations as a substrate for these principles, bridging “bio-inspired” and “brain-inspired” intelligence.
- 11:55-12:30
-
Invited Talk: Artificial selection of microbial communities - Wenying Shou Microbial communities perform functions beyond the reach of individual species, ranging from waste digestion to pathogen blockage to cheese fermentation. Community functions are often difficult to engineer, since they arise from complex and mostly unknown interactions among species. One possible approach to improving community function is through artificial community selection. This involves repeatedly selecting and propagating the most desirable communities, akin to classic breeding of livestock and crops. Although community selection is mechanism-agnostic, comparing evolved communities to their ancestors will reveal the mechanisms of resilience. In this talk, I will discuss challenges of community selection, and how to overcome them.
- 12:30-14:00
-
Lunch Break
- 14:00-14:35
-
Invited Talk: Information flow in self-organized developmental systems - David Brueckner Embryonic development relies on the ability of multi-cellular systems to collectively coordinate the self-organization of precise spatial patterns of cell fates. An inevitable obstacle to such coordination is intrinsic noise at the single-cell level, which constrains the amount of information accessible to cells for fate decisions. While the relevant molecular processes are increasingly well understood, we lack principled frameworks to quantify and predict how cells obtain sufficient information to reliably differentiate into the right fate at the right time and place. I will discuss how combining dynamical systems and information theory provides a mathematical language to analyze biological self-organization across diverse systems. Our approach can be used to define and measure the information content of observed patterns, to functionally assess the importance of various patterning mechanisms, and to predict optimal operating regimes of self-organizing systems. I will demonstrate how our framework reveals mechanisms of self-organization of in vitro stem cell systems in direct connection to experimental data, including intestinal organoids and gastruloids. This framework provides an avenue towards unifying the zoo of chemical and mechanical signaling processes that orchestrate embryonic development.
- 14:35-15:10
-
EPIC Member Talk: Cell assembly dynamics at boundaries in epithelial tissues - Daiki Umetsu Tissues are often subdivided into territories of unmixing cells during animal development. However, boundaries are challenged by continuous cell divisions and rearrangements. By combining live imaging and genetic or physical perturbations, we revealed both local increase in mechanical tension at boundaries and differential cell-cell adhesion maintain straight boundaries in a Drosophila epithelium. Our data also show that a gene encoding a Drosophila Toll-like receptor acts as an adhesion molecule to support the differential adhesion between neighbor cells. We will discuss how these mechanisms can be extrapolated to mechanisms by which cells with ectopic positional information are eliminated by surrounding cells.
- 15:10-15:20
-
Coffee Break
- 15:20-15:55
-
EPIC Member Talk: 3D Cell assemblies of motile cells in a non-metazoan system - Satoshi Sawai The cellular slime mold Dictyostelium discoideum is a member of the supergroup Amoebozoa – a large sister clade of fungi and animals. In this system, close to 100,000 cells migrate under starvation to form a fruiting body where cells undergo coordinated cellular movement and rearrangement. Our recent data obtained by light-sheet and confocal microscopy revealed how cell alignment reconfigures from the antero-posterior ordering to more convoluted three-dimensional layout with a diploblastic-like radial symmetry. The accompanying tissue elongation involves cell intercalation and amoeboid-to-epithelial like transitions. I will touch upon the essential cell-cell adhesion protein that we’ve identified as well as involvement of ancestral adhesion complex in this process. Our results hint at the toolbox necessary to facilitate evolution of morphogenetic motifs and the underlying cell behaviors.
- 15:55-16:30
-
Invited Talk: Computer simulations of the morphodynamics of dynamic cell assemblies - Tetsuya Hiraiwa Growing cell assemblies cultured in vitro, like organoids and epithelial cysts, provide powerful systems for studying the mechanisms determining the morphology of complex cellular structures, like organs. However, general mechanisms linking cellular behaviors to the emergent morphology of growing cell assemblies remain elusive. In the main part, we introduce our computational studies based on a multicellular phase-field model to investigate the morphodynamics of cell assemblies containing a fluid-filled cavity (lumen), grown from only a few cells, with a particular focus on lumen dynamics [Tanida 2026]. We also compare the results with experimental observations [Lu 2025; Lee 2026].
- 16:30-16:45
-
Coffee Break
- 16:45-18:30
-
Poster Session
- 10:00-10:35
-
EPIC Member Talk: - Kyogo Kawaguchi
- 10:35-11:10
-
Invited Talk: Synthetic RNA condensates and organelles - Lorenzo Di Michele Condensation of RNA and proteins forms functional membrane-less organelles that are central to cellular processes. Programming such condensation could enable metabolic engineering and the construction of synthetic cells. I will present a modular platform for engineering de novo RNA condensates from designed, branched RNA nanostructures that fold and assemble during transcription. Up to three orthogonal condensates can form simultaneously and selectively accumulate fluorophores via embedded fluorescent light-up aptamers. The condensates can be genetically encoded and expressed in synthetic cells, allowing control over number and size and enabling protein capture. Interactions can be tuned using linker RNAs to create biphasic condensates with controlled mixing. Finally, I will demonstrate deployment of the platform to engineering non-natural organelles in living E. coli.
- 11:10-11:20
-
Coffee Break
- 11:20-11:55
-
EPIC Member Talk: Nucleic-acid-based Smart Condensates for Molecular Computation - Masahiro Takinoue Synthetic liquid-like condensates formed via liquid-liquid phase separation (LLPS) are a key technology for developing synthetic cells and organelles. Although their thermodynamic stability has been well studied, controlling their dynamic properties remains challenging. This talk presents the dynamic control of programmable nucleic-acid-based condensates, where the sequences determine the stability and viscoelasticity. By coupling these condensates with enzymatic and strand displacement reactions, we achieved spatiotemporal control of LLPS condensates under nonequilibrium conditions. We demonstrate that these smart condensates can perform molecular computations, such as microRNA sensing, and exhibit complex mechanical behaviors. Synthetic LLPS condensate technologies will promote the development of functional synthetic cells and molecular robots.
- 11:55-12:30
-
Invited Talk: Physical organization of biomolecular condensates - Yongdae Shin Biomolecular condensates function in crowded cellular environments, yet how crowding shapes their composition remains unclear. In this talk, I will present an integrated approach using livecell reconstitution, biophysical perturbations, and computational modeling to examine how condensate density influences component behavior. Key concepts that will be discussed include the roles of sizebased exclusion, molecular interactions, and adaptive changes in condensate properties. These ideas offer broader insights into molecular flux and spatial organization in cells and provide a conceptual framework for how condensate density can govern molecular sorting.
- 12:30-14:00
-
Lunch Break
- 14:00-14:35
-
EPIC Member Talk: Evolutionary History of T1R Taste Receptors - Yasuka Toda In vertebrates, sweet and umami tastes are mediated by G protein–coupled receptors known as T1Rs. Humans possess only three T1R genes—T1R1, T1R2, and T1R3—which form heterodimeric receptors responsible for detecting umami and sweet stimuli. In our recent study, we expanded this view by identifying additional T1R members through comprehensive analyses of genome and transcriptome datasets from a wide range of jawed vertebrates. Our findings reveal that jawed vertebrates collectively harbor eleven distinct T1R genes. The long-term retention and diversification of these receptors appear to have been shaped by species-specific feeding ecologies. This work deepens our understanding of the evolutionary trajectories of T1R receptors and provides new insights into the origins of palatable taste modalities, namely umami and sweetness.
- 14:35-15:10
-
Invited Talk: Deciphering Olfaction with Structure, Evolutionary Genomics, and AI - Hiroaki Matsunami Our laboratory investigates the molecular mechanisms of olfaction through functional characterization of odorant receptors (ORs). ORs are remarkable for their large family size and rapid evolutionary diversification. We propose that this diversity is supported by the unique ability of olfactory sensory neurons to fold functional ORs despite cryptic mutations. To test this idea, we developed a consensus OR approach that corrects such mutations and overcomes a major expression barrier, enabling high-level heterologous expression. This strategy has facilitated atomic-level OR structure determination, providing new insights into odor binding and receptor activation.
- 15:10-15:20
-
Coffee Break
- 15:20-15:55
-
Invited Talk: From Signaling Complexity to Functional Principles in GPCR Biology - Ikuo Masuho GPCRs are central mediators of intercellular communication, detecting hormones and neurotransmitters and translating these cues into distinct cellular responses—such as gene expression, secretion, migration, or excitability—by engaging different G proteins. Using a universal live-cell platform, we quantified in real time the strength and speed of GPCR-driven G protein activation and defined receptor-specific signaling fingerprints. Applying this kinetic framework to more than 100 GPCRs uncovered general rules of coupling selectivity and offers a principled view of how GPCR signaling enables broad physiological control.
- 15:55-16:30
-
EPIC Member Talk: Dynamic Encoding of GPCR Signaling Revealed by Systematic Single-Cell Imaging - Kazuhiro Aoki G-protein-coupled receptors (GPCRs) translate diverse extracellular stimuli into intracellular responses through a limited set of conserved signaling proteins. How hundreds of receptors achieve signaling specificity remains unclear. Here, we systematically quantified the temporal dynamics of cAMP, Ca²⁺, RhoA, and ERK in response to activation of ~70 GPCRs at the single-cell level using live-cell imaging. Our dataset reveals striking diversity in signaling dynamics, including sustained, pulsatile, and transient patterns across receptors and pathways. Notably, the closely related noradrenergic receptors ADRA1A and ADRA1B generate sustained and oscillatory Ca²⁺ responses, respectively. Chimera analysis demonstrates that this difference is determined by the sixth and seventh transmembrane domains. These findings provide a systematic framework supporting dynamic encoding as a general principle of GPCR-mediated information processing and identify structural determinants that shape temporal signaling outputs.
- 16:30-16:35
-
Closing Remarks - Tetsuya Kobayashi
- 16:35-17:00
-
Open Discussion
- 17:00-
-
Reception
Participation is free of charge.
From young scientists to established leaders, all participants will gain valuable insights. Looking back, this gathering may well be remembered as a historic event.
Register nowAccess Information
Odakyu Line / Tokyo Metro-Chiyoda Line
8-min walk from Higashi-Kitazawa Station
12-min walk from Yoyogi Uehara Station
Keio Inokashira Line
10-min walk from Komaba-Todaimae Station
10-min walk from Ikenoue Station
Details
See "Access" page in Institute of Industrial Science, The University of Tokyo.
Map
- JSPS KAKENHI Grant-in-Aid for Transformative Research Areas (A) "EPIC Assembly: Emergence of Novel Functional Assembly by Evo-Physico Information Coupling" https://epic-assembly.crmind.net/
- Toyota Konpon Research Institute Investigative research project "Principles and Limitations of Frontier Exploration" https://konpon.toyota/exploration/



- JSPS KAKENHI Grant-in-Aid for Transformative Research Areas (B) "Life-Nonlife transition" https://life-nonlife.com
- JST CREST (JPMJCR2011) "Decoding of Information in Multi-Sourced Chemical Mixtures and its Applications"



